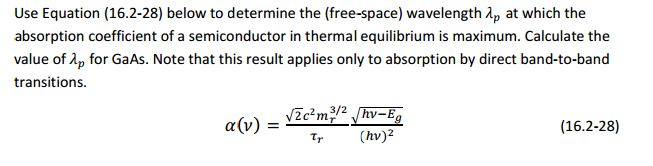
Absorption coefficients of silicon: A theoretical treatment: Journal of Applied Physics: Vol 123, No 18

Absorbance Measurements – the Quick Way to Determine Sample Concentration - Eppendorf Handling Solutions

Uncertainty analysis for the coefficient of band-to-band absorption of crystalline silicon: AIP Advances: Vol 5, No 6
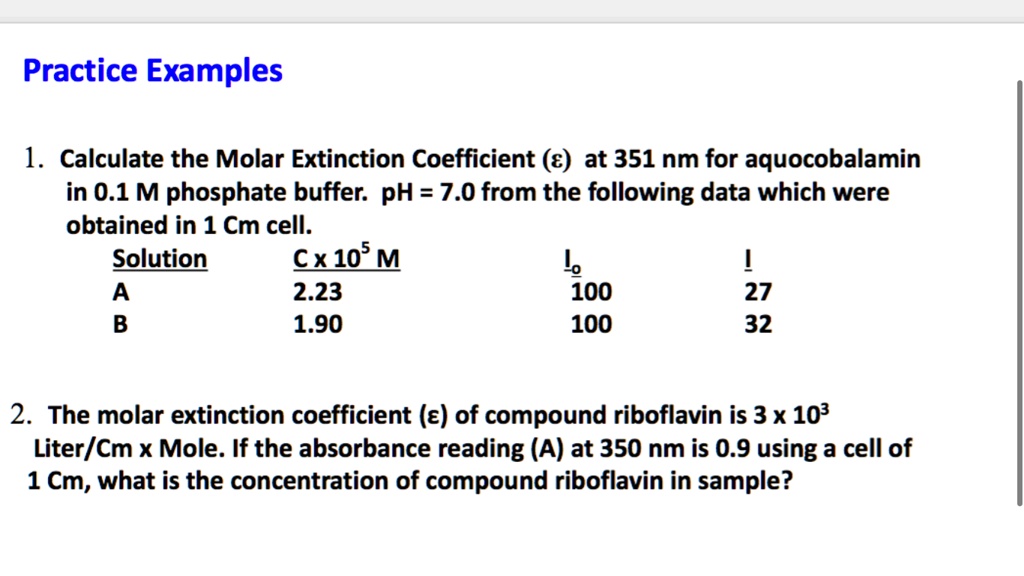

SOLVED: Practice Examples 1. Calculate the Molar Extinction Coefficient (€) at 351 nm for aquocobalamin in 0.1 M phosphate buffer: pH = 7.0 from the following data which were obtained in 1
![PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/af80a5d2beee749cff2f5d98dc4a8f0dfdd48074/2-Table1-1.png)
PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar

SOLVED: Practice Examples 1. Calculate the Molar Extinction Coefficient (€) at 351 nm for aquocobalamin in 0.1 M phosphate buffer: pH = 7.0 from the following data which were obtained in 1